Alphafold Training Optimization 01 | Data Processing Optimization
If there is one of the most exciting achievements in AI academia in 2021, then Alphafold deserves it.Alphafold2 achieved accuracy far beyond comparable models on the CASP14 protein prediction challenge, and for the first time improved the accuracy of protein structure prediction to the atomic level-which already close to the level of experimental measurements.
The Phantom AI team successfully ran Alphafold2 training on the Firefly II platform shortly after Alphafold2's launch, as detailed in our previous articleFirefly Running Model | Alphafold Protein Structure Prediction.. However, the complexity of Alphafold2's own model and the flaws of the open source code, and the special training platform of Phantom Cube AI, made the reproduction resource utilization very inefficient at the beginning, and the training cost unacceptable.
In order to improve the training efficiency of Alphafold and help researchers and developers to lower the research threshold, the Phantom AI team has made a number of optimizations to the Alphafold2 model, which have dramatically improved the efficiency of model training. Our work can be summarized in the following three points:
- Refactor the code to solve the problem of not being able to train on multiple machines and cards in the open-source version, and utilize data parallelism to dramatically improve Alphafold2 training efficiency and reduce the reproduction threshold;
- Optimize the data processing logic and dramatically increase GPU utilization from around 40% to over 90%;
- Using hfai high performance tools to transform the model, Alphafold2 is deeply integrated into the training platform of Phantom Cube AI to significantly improve training performance.
This post starts with a conversation about how to optimize the processing of protein data for efficient training performance.
model store:https://github.com/HFAiLab/alphafold-optimized
Input Optimization
Alphafold's input features can be categorized into three such categories:
- The protein sequence itself
- Multi-Sequence Alignment (MSA) feature
- Template Protein Template Features
They vary so much between different protein sequence lengths that the time required for processing can range from a few seconds to thousands of seconds, which can greatly affect the efficiency of model training. For example, in a 20-minute training process, the processing of a single long sample may block the overall training for 10 minutes. In a multi-computer, multi-card, large batch scenario, it is almost inevitable that the processing time of some data within each batch is extremely long.
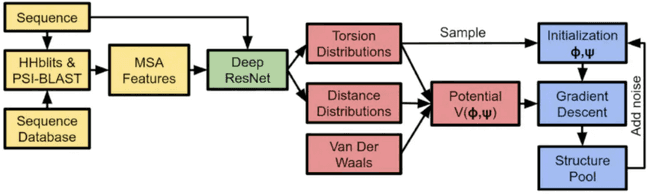
As shown in the above figure, in the initial stage of training Alphafold, the input features are derived from sequence database search, computed by biological tools (JackHmmer/HHSearch/...), which need to rely on a large amount of CPU computing power support. Throughout the process of Alphafold model training, CPU arithmetic power often becomes an important bottleneck that hinders the performance improvement of Alphafold training.
To address these issues, Phantom AI usesFeature preprocessing与Feature CutsTwo schemes optimize the inputs. Briefly, feature preprocessing utilizes a dedicated CPU cluster to pre-do a large number of database search tasks, biological tool computation tasks to get the feature set and candidate set; in the model training phase, the model directly pulls the feature set and candidate set, screens the features, and appropriately crops the features of the long protein sequences. By this method, we realized a 4-5 times improvement in the training performance of alphafold, which will be described in detail in the next section.
1. Feature pre-processing
In a typical model training task, the bulk of the time consumption lies in the GPU's forward inference and backpropagation computations, and the data processing can be masked by asynchronous Dataloader loading to cover the time overhead. However, in Alphafold's training, one iteration of the model over the processed data takes about 11 seconds (based on the A100 GPU), while the preprocessing of this one data takes about 150 seconds (on an entire EPYC 64-core processor), which is 13 times longer than the GPU computation time, and cannot be hidden by GPU computation at all. In this case.The theoretical upper limit of GPU utilization is also only 7.31 TP3T, which is very inefficient. At the same time, each machine on the Firefly II cluster is equipped with 8 GPUs and one CPU, according to the original multi-card parallel training method, CPU resources will be even more scarce, so optimizing feature processing during training is meaningless, and we need to find other alternatives.
We chooseSeparate the feature processing separately, use CPU cluster arithmetic to pre-complete a part of the task, and save the obtained candidate sample data. In order to improve the efficiency of the service and maximize CPU utilization, we used Python multiprocessing to speed up feature preprocessing in parallel. However, feature processing involves several different biological tools, and each tool has different CPU utilization, so direct parallelization can easily lead to too many processes, which generates a large CPU scheduling preemption overhead. Statistically, the CPU resource usage during the processing of several different types of features is shown in the table below:
| Protein datasets | Processing tools | Time consumed (seconds) | CPU utilization | Total computing power requirements (minutes/core) |
|---|---|---|---|---|
| Uniref90 | JackHmmer | 567 | 4-core | 37.80 |
| Magnify | JackHmmer | 851 | 4-core | 56.73 |
| Bfd-small | JackHmmer | 308 | 4-core | 20.53 |
| Pdb70 | HHSearch | 62 | 64-core | 66.13 |
From the above table, we can observe that: the first three datasets use JackHmmer, which basically can only use 4-core CPU resources; while Pdb70 uses HHSearch, which is able to use all the resources on the CPU. Therefore, weNeed to create different process pools for different processing tools for scheduling allocation. The optimization strategy we use is to launch multiple JackHmmer tasks at the same time and launch an HHSearch task at regular intervals.
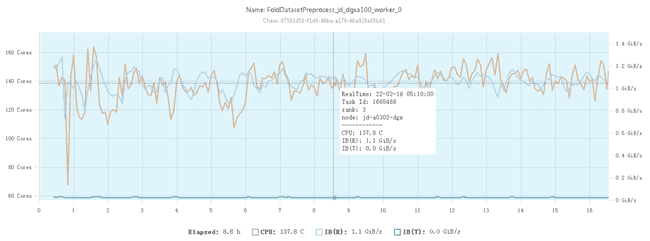
The CPU occupancy of the optimized feature preprocessing on a 128-core CPU node is shown below:
It can be seen that at this point it is already possible to use CPU resources basically at a relatively stable 100% utilization rate.
it's worth noting that...For MSA features and Template features, Alphafold's paper mentions that the training is a search process to randomly select the most suitable 4 features as inputs, so the preprocessing can provide a suitable candidate set, while the final model training which 4 features to select as inputs cannot be replaced by the preprocessing part. features as input when the final model is trained, the preprocessing part cannot be replaced.
The intermediate results from the pre-processing of the above features can be obtained by using Phantom AI's self-developedFFRecordformat for storage to take full advantage of the Firefly cluster's high-speed file system, 3FS, as detailed in the previous articlePhantom Force | High Speed File System: 3FS。
2. Feature cropping
As mentioned earlier, the processing of protein sequences of different lengths may cause training to be blocked, and in addition to switching runtime processing to preprocessing, we can alsoCropping of longer features. However, it should be noted that feature cropping of features will inevitably affect the effectiveness of model training, so a balance needs to be adjusted between model performance and training efficiency.
Feature cropping is only for the processing of MSA features and Template features, because its main goal is to find more similar features (sequence, structure) to enhance the expression ability of the model. Feature cropping can effectively reduce the feature processing overhead for complex proteins and improve the overall training efficiency. Protein sequence itself features are not suitable for feature cropping.
2.1 MSA feature cropping
MSA features are a set of other biological protein sequences that are similar to the input human protein sequence and are used to help the model Encoder part to form the feature vector of the protein. If the length of the input protein sequence is N amino acids and the number of homologous proteins found in the preprocessing stage is M, the size of the MSA features to be preprocessed is NxM. It can be seen that under the premise that protein sequences cannot be cropped, the time consuming part can only be reduced by cropping the number of homologous proteins, i.e., by discarding a part of the homologous sequences that were previously found.
Protein sequence length N, number of MSA feature sequences M, and final processing time T. They satisfy the following linear relationship: T=C×N×MT=C×N×M
C is a fixed constant and is related to the processing power of the machine. In our experiments, we found that the maximum length of time to make training unblocked was 150 seconds, so we can derive the maximum number of MSA features per input protein sequence based on the formula above. In practice, in our experiments, less than 101 TP3T of sequences needed to be cropped for MSA features, so this can greatly improve the efficiency of the training without compromising performance as much as possible.
2.2 Template Feature Cropping
Template features are designed to extract protein sequences that are similar in spatial structure to the protein to be predicted, and are used to help the Structure Module part of the model to form a feature representation that outputs the spatial structure of the protein. Similarly, there may be hundreds of templates for a single protein, and although only 4 templates need to be fed into the model to participate in the training of Alphafold eventually, not all of them may actually match the protein. Thus once a situation is encountered where a large number of templates do not match, the process of checking the template sequence may take up to ten minutes.
The processing of Template features can be accelerated in parallel, but due to the problems with the PyTorch Dataloader daemon and the limitation of the amount of CPU resources, this approach is not very applicable in real training. Therefore, we also consider cropping Template features to reduce the time consumed in this step.
- Feature preprocessing to obtain Template feature candidate set, sorted by match;
- Try to use a template that better matches the original protein when training;
- There is a fixed time limit for the template matching process, and the timeout does the cropping.
Later experiments will demonstrate the effectiveness of the above methods in actual training.
test
First, we start by testing Alphafold's time overhead per Step for training on a single A100 graphics card as a benchmark:
| Stage | Mean duration (s) | Percentage (%) |
|---|---|---|
| Training Step | 12.65 | 100 |
| Get Training Batch | 1.87 | 14.78 |
| Forward | 4.24 | 33.52 |
| Zero Grad | 0.17 | 1.31 |
| Exponential Moving Average | 0.14 | 1.0 |
| Optimizer Step | 0.45 | 3.42 |
| Backward | 5.93 | 46.88 |
From the table, we can see that it takes 12.65 seconds to train one Step, where the most important time overhead lies in the Backward, Forward and Get Training Batch steps. Due to the small amount of data used and the lack of multi-card parallelism, the performance bottleneck caused by feature processing and the overhead of DDP have not yet been realized.
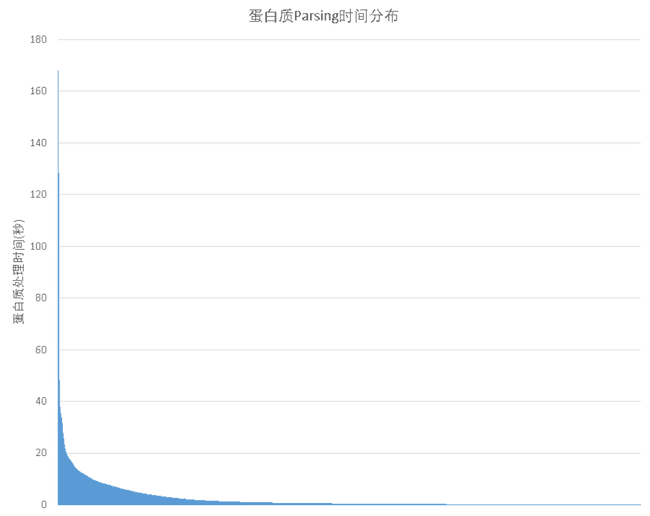
And when there are more training data, the difference in processing time between different training data becomes especially prominent. As shown in the figure below, most of the input data takes only a few seconds to process, however a small number of proteins will require hundreds of seconds of preprocessing time in extreme cases.
Thus once one such input data is encountered, it will cause the whole training to block. And assuming that only 51 TP3T out of all the training data will lead to training blocking, the probability of each batch being blocked when training Alphafold (the original paper parameter) using a batch_size of 128 would be.
Pstuck=1-(1-0.05)128=99.86%Pstuck=1−(1−0.05)128=99.86%
This is a blocking situation that is encountered in almost every step of the training. At this point, the GPU utilization of the entire training will drop to less than 20% in extreme cases, with most of the time spent waiting for one of the batch to finish processing the data.
For example, if we use 16 nodes and 128 A100s for training without optimization, the time spent in the first 4 steps is 413s, 177s, 100s, 101s, and the GPU utilization in each step is less than 10%. It can be seen that if we don't optimize the data processing at this time, it is difficult to train normally.
1. Feature preprocessing tests
We randomly selected 35,000 protein sequences for parsing out of a total of 110,000 protein sequence training data and counted the time consumed by these sequences using direct training-time processing and feature preprocessing approaches, respectively:
| carbohydrate | Original processing time (s) | Time after preprocessing using features (s) |
|---|---|---|
| 3j3q | 145.9 | 46.6 |
| 3j3y | 129.8 | 41.0 |
| 6u42 | 78.9 | 27.5 |
As shown in the table above, for sequences that take longer than 10 seconds to process, the feature parsing process, which originally took 150 seconds, only takes about 50 seconds after optimization, which greatly suppresses the blocking effect of very long sequences on the overall training. Even for small sequences, a speedup of nearly 3 times is obtained.
Next, let's see how much benefit using the feature preprocessing results can bring to the GPU utilization during training and the total training time. Here, we compare the performance gains after the feature preprocessing method without feature cropping first. The results are shown in the table below:
| No feature preprocessing | Perform feature preprocessing | |
|---|---|---|
| Length of data processing at the beginning of each round (s) | 451 | 393 |
| Average time per step (s) | 50.3 | 23.57 |
| Number of times training was blocked | 10 | 6 |
| GPU Utilization | 19.58% | 42.37% |
It can be seen that performing feature preprocessing can obtain a very significant performance improvement, with GPU utilization doubled. However, at the same time, too much training blocking causes the overall GPU utilization to remain unsatisfactory.
2. MSA Feature Cropping Test
Next, we test the gain of using MSA feature cropping on GPU utilization and total training time during training. Here the MSA feature cropping time constant is 2e-6, no Template feature cropping is performed, and feature preprocessing is used for all. The results are shown in the following table:
| No MSA cuts | Cutting MSA Features | |
|---|---|---|
| Length of data processing at the beginning of each round (s) | 393 | 200 |
| Average time per step (s) | 23.57 | 10.87 |
| Number of times training was blocked | 6 | 1 |
| GPU Utilization | 42.37% | 90.2% |
It can be seen that GPU utilization can be more than doubled again after using MSA feature cropping, and training blocking occurs only once. This method can effectively improve the training efficiency.
3. Template feature crop test
After that, we then test the effect of Template feature cropping on training efficiency. The time limit is 150 seconds, both using feature preprocessing without MSA feature processing. The results are shown in the following table:
| No Template Cropping | Cutting Template Features | |
|---|---|---|
| Length of data processing at the beginning of each round (s) | 393 | 201 |
| Average time per step (s) | 23.57 | 11.65 |
| Number of times training was blocked | 6 | 1 |
| GPU Utilization | 42.37% | 90.2% |
GPU utilization is also more than doubled with Template feature cropping.
wrap-up
As one of the most influential work in the field of AI in 2021, the reproduction of Alphafold is undoubtedly meaningful and can help more researchers and developers to explore the prospect of this direction. By optimizing the data input, we can achieve a GPU usage of more than 90%, which efficiently utilizes the cluster computing resources. At the same time, Phantom AI also has many efficient training tools, such as hfreduce, 3FS, operator and so on, can they make Alphafold's training further accelerate? The next installment of the technology sharing will be presented to you.
- 本文作者: suopu
您可以转载、不违背作品原意地摘录及引用本技术博客的内容,但必须遵守以下条款: 署名 — 您应当署名原作者,但不得以任何方式暗示幻方为您背书,亦不会对幻方的权利造成任何负面影响。 非商业性使用 — 您不得将本技术博客内容用于商业目的。 禁止演绎 — 如果基于该内容改编、转换、或者再创作,您不得公开或分发被修改内容,该内容仅可供个人使用。